© Borgis - Postępy Nauk Medycznych 1/2015, s. 19-24
*Cezary Cybulski1, Aniruddh Kashyap1, Wojciech Kluźniak1, Dominika Wokołorczyk1, Adam Gołąb2, Marcin Słojewski2, Bartosz Gliniewicz3, Andrzej Antczak4, Andrzej Sikorski2, Jan Lubiński1
Dziedziczny rak prostaty
Hereditary prostate cancer
1Department of Genetics and Pathology, International Hereditary Cancer Center, Pomeranian Medical University, Szczecin
Head of Department: prof. Jan Lubiński, MD, PhD
2Clinic of Urology, Pomeranian Medical University, Szczecin
Head of Clinic: Marcin Słojewski, MD, PhD
3Division of Urology, Maria Skłodowska-Curie Hospital, Szczecin
Head of Division: Jerzy Świtała, MD, PhD
4Department and Clinic of Urology and Urologic Oncology, University of Medical Sciences, Poznań
Head of Department and Clinic: Andrzej Antczak, MD, PhD
Streszczenie
Badania epidemiologiczne ostatniego dwudziestolecia dowodzą, że czynniki genetyczne mają ogromne znaczenie w etiologii raka gruczołu krokowego, jednakże geny związane z dziedziczną predyspozycją do tego nowotworu pozostają w dużej mierze niepoznane. W Polsce prowadzimy badania, których celem jest wyjaśnienie podłoża genetycznego raka prostaty. Dotychczas wykryliśmy jednoznaczny związek pomiędzy nosicielstwem zmian konstytucyjnych genów CHEK2, NBS1, HOXB13 oraz zmiany polimorficznej w regionie 8q24 (rs188140481) a zwiększonym ryzykiem zachorowania na raka prostaty w polskiej populacji. Nasze badania dowodzą, że do poznanych genetycznych markerów wysokiego ryzyka raka prostaty również można zaliczyć nosicielstwo tych specyficznych zmian DNA, u których w rodzinie stwierdzono co najmniej jedno zachorowanie na raka prostaty u krewnego I lub II stopnia (ryzyko zachorowania zwiększone około 3-8-krotnie). W oparciu o wyniki naszych badań uważamy, że badanie genetycznej predyspozycji do raka prostaty w polskiej populacji może być oparte o wykrywanie siedmiu konstytucyjnych zmian DNA (IVS2+1G>A, 1100delC, del5395, I157T in CHEK2; 657del5 in NBS1; G84E in HOXB13; allele A of rs188140481). Identyfikacja genetycznych markerów podatności na raka prostaty ma na celu usprawnienie profilaktyki, diagnostyki i postępowania z rakiem stercza w Polsce.
Summary
Epidemiologic research conducted over the last two decades has led us to believe that inherited factors play an important role in the etiology of prostate cancer, but the genes, which underlie the inherited susceptibility are elusive. In Poland, we have initiated a program to identify DNA variants, which confer an increased risk of prostate cancer. We found that germline mutations in CHEK2, NBS1, HOXB13 and the rs188140481 variant in the 8q24 region confer an increased prostate cancer risk in Polish men. Our studies provide evidence that the list of known genetic markers of high risk of prostate cancer can be extended by these specific variants in men with a positive family history of prostate cancer in at least one first or second degree relative (the risk increased about 3-8 fold). Based on our findings, we recommend that genetic testing for prostate cancer susceptibility in Poland be based on the seven founder alleles (IVS2 + 1G > A, 1100delC, del5395, I157T in CHEK2; 657del5 in NBS1; G84E in HOXB13; allele A of rs188140481). Identification of genetic markers of prostate cancer susceptibility will improve prevention, diagnosis and management of prostate cancer in Poland.

Familial clustering of prostate cancer was first described in 1955, and a term hereditary prostate cancer (HPC) was first used in 1992 by Carter who reported the results of linkage analysis in a series of 691 men with prostate cancer (PC) (1). This analysis revealed that 9% cases of familial clustering of prostate cancer is associated with a single rare allele. Penetrance of this allele was 88% to age of 85. The allele conferring high risk for prostate cancer was found on long arm of chromosome 1 (1q24-25), and this loci was named HPC1 (2).
Since that time, a number of prostate susceptibility chromosomal loci have been identified, but no particular genes at these loci were found to be associated with a significant proportion of HPC and clinical importance of these loci is still very limited. Although the molecular basic of HPC largely remains unknown, epidemiologic research clearly shows that inherited factors play an important role in the etiology of prostate cancer. Genes of high penetrance may be responsible for 5-10% of prostate cancer cases and for as many as 30-40% of early onset prostate cancer (3). Scandinavian study of twins suggested that the heritability of prostate cancer may be as high as 42% (4).
Family history and prostate cancer risk
Familial clustering of prostate cancer is the most important risk factor for prostate cancer (3, 5). Many studies show an elevated risk of PC in brothers and sons of affected individuals. It is important to note that the risk of PC (especially early onset) is higher in relatives of individuals affected by prostate cancer in early age. Some of the studies show that the risk is higher in brother than in sons of PC patients (it is possible that inheritance is X-linked or HPC could be an autosomal recessive trait in some families). The risk of prostate cancer by the presence of prostate cancer in first and second degree relatives is shown in table 1.
Table 1. Family history and prostate cancer risk.
| Family history | Relative Risk |
| Negative | 1 |
| Father with PC at age of 60 or above | 1.5 |
| 1 brother with PC at age of 60 or above | 2 |
| Father with PC before age of 60 | 2.5 |
| 1 brother with PC before age of 60 | 3 |
| 2 first degree relatives with PC | 4 |
| 3 or more relatives with PC | 5 |
Clinical criteria for HPC
Clinical criteria for HPC (fig. 1a and 1b) (6):
1. Definitive diagnosis of HPC:
a) PC in 3 or more first degree relatives; or
b) PC in 3 generations; or
c) PC at age below 56, in two or more relatives.
2. Diagnosis of cases suspected for HPC:
a) PC in 3 or more relatives but does not fulfil criterion a) or b) for definitive diagnosis; or
b) PC in 2 relatives, including at least one diagnosed below age of 60 and/or vertical transmission but without fulfilling criterion c) for definitive diagnosis; or
c) at least one PC below age of 50, not matching criteria for definitive diagnosis.
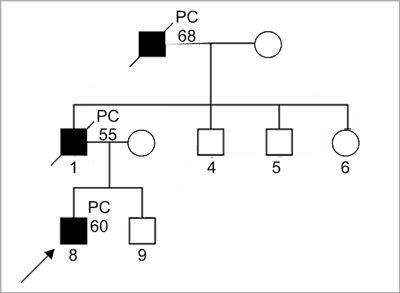
Fig. 1a. Pedigree of family with diagnosis of HPC.
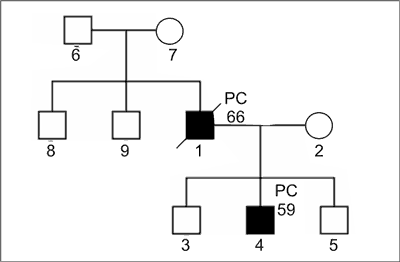
Fig. 1b. Pedigree of a family suspected for HPC, which does not fulfil clinical criteria.
Clinical characteristics of HPC
The most important features of HPC include: autosomal dominant inheritance of prostate cancer (rarely autosomal recessive or X-linked dominant inheritance) and early age of diagnosis – the mean age below 56, so 6-7 years younger than that seen in sporadic cases (2). Because of the early onset of disease, a bigger proportion of HPC patients die (75%), than in sporadic cases (50%) (7, 8).
Other cancers in families with HPC
Epidemiological studies provide evidence that in families with prostate cancer the risk of this cancer is increased, but the data are controversial regarding increased risk of other cancers. Some studies suggest that risk for brain tumours, stomach cancer or breast cancer is increased in HPC, but in most studies HPC is characterized as susceptibility to prostate cancer in isolation (site specific syndrome). The controversy will remain questioned until the majority of HPC susceptibility genes are identified (9).
Hereditary cancer syndromes and PC risk
Mutations in BRCA1 or BRCA2 gene confer high risk of breast and ovarian cancer. Several studies suggested two-fold increased risk of prostate cancer in Ashkenazi Jewish men with a BRCA1 mutation (185delAG or 5382insC) (10-12). While studies in other non-Jewish populations have found little or no evidence of an increased risk for prostate cancer in BRCA1 mutation carriers (13-16). The reason of this difference may be a different spectrum of mutations in BRCA1 gene and/or other modifying factors in particular populations.
Association of BRCA2 mutations with increased prostate cancer risk is well documented. It was reported that, carriers of germ-line mutations in BRCA2 are at 5 fold increased risk of PC, the risk higher (increased by 7 fold) to age of 65, and even 20-fold to age of 56 (9). Recent studies suggest that BRCA2 carriers are more likely to develop aggressive prostate cancer (high grading G3 and G4) in early age (on an average 5 years younger than non-carriers). Prognosis was reported to be much worse for men with a BRCA2 mutation, i.a. one study showed that the mean survival was approximatelly 2 years in BRCA2 carriers versus 12 years in non-carriers. However, mutations in BRCA2 or BRCA1 are rare and the contribution of these two genes to HPC is relatively small, which is largely a site specific syndrome.
In addition increased prevalence of prostate cancer was observed in families with Cowden syndrome, Li-Fraumeni syndrome, and hereditary stomach cancer caused by mutations of E-cadherin (3, 9).
Prostate cancer susceptibility genes (HPC candidate genes)
Through linkage analysis, numerous prostate cancer susceptibility chromosomal loci have been identified (i.e. HPC1 (1q24-25), PCaP (1q42-43), HPCX (Xq27-28), CAPB (1p36), HPC2 (17p12) and HPC20 (20q13)). From these regions three candidate susceptibility genes (i.a. RNASEL and MSR1) have been positionally cloned. Germ-line mutations in RNASEL and MSR1 genes have been reported in several families affected with hereditary prostate cancer in USA, and frequent polymorphisms of these genes were reported as variants of low penetrance for prostate cancer (17-19). Unfortunately further studies provided evidence that none of these genes found for HPC is a high-risk susceptibility gene for prostate cancer, including studies in Polish population (20).
HOXB13 gene
Through linkage analyses in HPC families a number of chromosomal loci was found to be associated with PC. The first gene conferring high risk of PC identified in one of these loci was only discovered in 2012. Ewing et al. performed next generation sequencing of chromosomal region 17q21-22 (above 200 genes) in 94 families with PC and found a recurrent mutation G84E in HOXB13 gene (21). The mutation was confirmed in familial segregation analysis and association analysis showed high risk of prostate cancer linked to this mutation (OR = 20; p < 0.0001). High risk of prostate cancer in carriers of mutation G84E in HOXB13 gene conferred in different studies and in different populations, including Polish population (22).
CHEK2 gene
Powyżej zamieściliśmy fragment artykułu, do którego możesz uzyskać pełny dostęp.
Mam kod dostępu
- Aby uzyskać płatny dostęp do pełnej treści powyższego artykułu albo wszystkich artykułów (w zależności od wybranej opcji), należy wprowadzić kod.
- Wprowadzając kod, akceptują Państwo treść Regulaminu oraz potwierdzają zapoznanie się z nim.
- Aby kupić kod proszę skorzystać z jednej z poniższych opcji.
Opcja #1
29 zł
Wybieram
- dostęp do tego artykułu
- dostęp na 7 dni
uzyskany kod musi być wprowadzony na stronie artykułu, do którego został wykupiony
Opcja #2
69 zł
Wybieram
- dostęp do tego i pozostałych ponad 7000 artykułów
- dostęp na 30 dni
- najpopularniejsza opcja
Opcja #3
129 zł
Wybieram
- dostęp do tego i pozostałych ponad 7000 artykułów
- dostęp na 90 dni
- oszczędzasz 78 zł
Piśmiennictwo
1. Carter BS, Beaty TH, Steinberg GD et al.: Mendelian inheritance of familial prostate cancer. Proc Natl Acad Sci USA 1992; 89: 3367-3371.
2. Smith JR, Freije D, Carpten JD et al.: Major susceptibility locus for prostate cancer on chromosome 1 suggested by a genome-wide search. Science 1996; 274: 1371-1374.
3. Bratt O: Hereditary prostate cancer: clinical aspects. J Urol 2002; 168: 906-913.
4. Lichtenstein P, Holm NV, Verkasalo PK et al.: Environmental and heritable factors in the causation of cancer-analyses of cohorts of twins from Sweden, Denmark, and Finland. N Engl J Med 2000; 343: 78-85.
5. Virtanen A, Gomari M, Kranse R, Stenman UH: Estimation of prostate cancer probability by logistic regression: free and total prostate-specific antigen, digital rectal examination, and heredity are significant variables. Clin Chem 1999; 45: 987-994.
6. Carter BS, Bova GS, Beaty TH et al.: Hereditary prostate cancer: epidemiologic and clinical features. J Urol 1993; 150: 797-802.
7. Bratt O, Damber JE, Emanuelsson M, Gronberg H: Hereditary prostate cancer: clinical characteristics and survival. J Urol 2002; 167: 2423-2426.
8. Keetch DW, Humphrey PA, Smith DS et al.: Clinical and pathological features of hereditary prostate cancer. J Urol 1996; 155: 1841-1843.
9. Sigurdsson S, Thorlacius S, Tomasson J et al.: BRCA2 mutation in Icelandic prostate cancer patients. J Mol Med 1997; 75: 758-761.
10. Struewing JP, Hartge P, Wacholder S et al.: The risk of cancer associated with specific mutations of BRCA1 and BRCA2 among Ashkenazi Jews. N Engl J Med 1997; 336: 1401-1408.
11. Warner E, Foulkes W, Goodwin P et al.: Prevalence and penetrance of BRCA1 and BRCA2 gene mutations in unselected Ashkenazi Jewish women with breast cancer. J Natl Cancer Inst 1999; 91: 1241-1217.
12. Giusti RM, Rutter JL, Duray PH et al.: A twofold increase in BRCA mutation related prostate cancer among Ashkenazi Israelis is not associated with distinctive histopathology. J Med Genet 2003; 40: 787-792.
13. Thompson D, Easton DF: Breast Cancer Linkage Consortium: Cancer incidence in BRCA1 mutation carriers. J Natl Cancer Inst 2002; 94: 1358-1365.
14. Sinclair CS, Berry R, Schaid D et al.: BRCA1 and BRCA2 have a limited role in familial prostate cancer. Cancer Res 2000; 60: 1371-1375.
15. Ikonen T, Matikainen MP, Syrjäkoski K et al.: BRCA1 and BRCA2 mutations have no major role in predisposition to prostate cancer in Finland. J Med Genet 2003; 40: E98.
16. Zuhlke KA, Madeoy JJ, Beebe-Dimmer J et al.: Truncating BRCA1 mutations are uncommon in a cohort of hereditary prostate cancer families with evidence of linkage to 17q markers. Clin Cancer Res 2004; 10: 5975-5980.
17. Carpten J, Nupponen N, Isaacs S et al.: Germline mutations in the ribonuclease L gene in families showing linkage with HPC1. Nat Genet 2002; 30: 181-184.
18. Casey G, Neville PJ, Plummer SJ et al.: RNASEL Arg462Gln variant is implicated in up to 13% of prostate cancer cases. Nat Genet 2002; 32: 581-583.
19. Xu J, Zheng SL, Komiya A et al.: Germline mutations and sequence variants of the macrophage scavenger receptor 1 gene are associated with prostate cancer risk. Nat Genet 2002; 32: 321-325.
20. Cybulski C, Wokołorczyk D, Jakubowska A et al.: DNA variation in MSR1, RNASEL and e-cadherin genes and prostate cancer in Poland. Urol Int 2007; 79: 44-49.
21. Ewing CM, Ray AM, Lange EM et al.: Germline mutations in HOXB13 and prostate-cancer risk. N Engl J Med 2012; 366(2): 141-149.
22. Kluźniak W, Wokołorczyk D, Gołąb A et al.: The G84E mutation in the HOXB13 gene is associated with an increased risk of prostate cancer in Poland. Prostate 2013; 73: 542-548.
23. Dong X, Wang L, Taniguchi K et al.: Mutations in CHEK2 associated with prostate cancer risk. Am J Hum Genet 2003; 72: 270-280.
24. Seppälä EH, Ikonen T, Mononen N et al.: CHEK2 variants associate with hereditary prostate cancer. Br J Cancer 2003; 89: 1966-1970.
25. Cybulski C, Górski B, Debniak T et al.: NBS1 is a prostate cancer susceptibility gene. Cancer Res 2004; 64: 1215-1219.
26. Cybulski C, Wokołorczyk D, Kluźniak W et al.: An Inherited NBS1 Mutation is Associated with Poor Prognosis Prostate Cancer. Br J Cancer 2013; 108: 461-468.
27. Kibel AS, Suarez BK, Belani J, Oh J: CDKN1A and CDKN1B polymorphisms and risk of advanced prostate carcinoma. Cancer Res 2003; 63: 2033-2036.
28. Angèle S, Falconer A, Edwards SM et al.: ATM polymorphisms as risk factors for prostate cancer development. Br J Cancer 2004; 91: 783-787.
29. Rybicki BA, Conti DV, Moreira A et al.: DNA repair gene XRCC1 and XPD polymorphisms and risk of prostate cancer. Cancer Epidemiol Biomarkers Prev 2004; 13: 23-29.
30. Eeles RA, Olama AA, Benlloch S et al.: Identification of 23 new prostate cancer susceptibility loci using the iCOGS custom genotyping array. Nat Genet 2013; 45: 385-391.
31. Cybulski C, Huzarski T, Górski B et al.: A novel founder CHEK2 mutation is associated with increased prostate cancer risk. Cancer Res 2004; 64: 2677-2679.
32. Cybulski C, Wokołorczyk D, Huzarski T et al.: A large germline deletion in the CHEK2 kinase gene is associated with an increased risk of prostate cancer. J Med Genet 2006; 43: 863-866.
33. Cybulski C, Wokołorczyk D, Kluźniak W et al.: A personalised approach to prostate cancer screening based on genotyping of risk founder alleles. Br J Cancer 2013; 108: 2601-2609.
34. von Eschenbach A, Ho R, Murphy GP et al.: American Cancer Society guidelines for the early detection of prostate cancer: update, June 10, 1997. Cancer 1997; 80: 1805-1807.
35. Machoy P, Lubiński J: Dziedziczny rak prostaty. Urologia Polska 2002; 55: 3.
36. Gudmundsson J, Sulem P, Gudbjartsson DF et al.: A study based on whole-genome sequencing yields a rare variant at 8q24 associated with prostate cancer. Nat Genet 2012; 44: 1326-1329.
37. Antczak A, Wokołorczyk D, Kluźniak W et al.: The variant allele of the rs188140481 polymorphism confers a moderate increase in the risk of prostate cancer in Polish men. Eur J Cancer Prev 2014. In press.
38. Ilic D, Neuberger MM, Djulbegovic M, Dahm P: Screening for prostate cancer. Cochrane Database Syst Rev 1: CD004720. doi: 10.1002/14651858.
39. Smith RA, Cokkinides V, Brawley OW: Cancer screening in the United States, 2009: a review of current American Cancer Society guidelines and issues in cancer screening. CA Cancer J Clin 2009; 59: 27-41.


